|
Biomedical engineering is expensive, really expensive. Hundreds of reagents, plates, pipettes, and pieces of equipment populate a lab. It is not a cheap affair, which means it is important for researchers to save money where they can. One of the easiest ways to do this is to use small amounts of expensive compounds.
In the laboratory where I work, the proteins we isolate are the products of many labor and resource-intensive steps, and are thus very valuable. To isolate a single protein sample, DNA purification reagents, Lipofectamine, lysing agents, and countless flasks must be expended. As such, it is in the lab’s best interest to conserve these proteins. This can be done by knowing the concentration of protein in each sample, which allows researchers to calculate the minimum volume that must be used for their tests. In order to find the concentration of the proteins, a Bradford protein assay is used. To conduct this assay, very small amounts of samples (as low as 5µL) are added to wells. A combination of reagents is then added to the wells, and a reaction begins which initiates a chromatic change. When there is no protein in the well, the solution trends towards clear. When there is an abundance of protein in the well, a dark blue color becomes apparent. In order to quantify the results of this assay, it is important to use a series of standards (solutions with known protein concentrations) as controls. A spectrophotometer then takes the absorbance of each well. Beer’s law states that absorbance and concentration are directly related, so a calibration curve can be created from the standards. Each sample’s absorbance is then run against the calibration curve in order to have its concentration extrapolated. This process may seem lengthy, but it can be done in around an hour with the proper equipment. A Bradford assay helps researchers to stay efficient in their use of protein samples, with one practical application being helping to determine the amount of sample required to conduct a Western blot. Thanks to the Bradford assay and tests like it, good research can also be efficient research.
0 Comments
Today, I want to talk a little bit about how genetic engineers can assess the success of their work. There is no point in investing time and money into developing a genetically modified organism if scientists cannot ascertain how their efforts are panning out. One specific technique that researchers can use to accomplish this is the “western blot.”
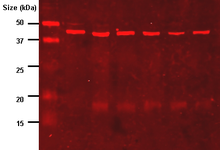
The central dogma of microbiology is the idea that physiological traits are the product of proteins, which themselves are the result of genes being expressed. Physical traits are easy to see in organisms which exist on the macro scale, but when scientists are dealing with entities such as microscopic cell colonies and monolayers, it can get very difficult to see the effects that a gene is having. It is often easier to check for the presence of the protein which creates a trait, which is where the western blot comes in. Western blotting’s first step utilizes gel electrophoresis in order to separate proteins within a sample on the basis of size or charge. A sample is first processed in order to lyse the cells and expose the proteins, and a series of buffers and detergents can be used in order to protect said proteins. This sample is then placed in a well on a gel electrophoresis rig in order for the proteins to separate. As a control, a solution of proteins with known weights is normally run in the same gel, this provides a sort of “ruler” that lets researchers determine the size of their proteins with greater accuracy (the control also tells researchers when the gel has finished running). The proteins are denatured in order to leave only their primary structure and are separated. The larger proteins have a harder time making their way through the gel, so they do not travel as far as the smaller proteins. After the gel is finished, it is time to visualize the results. The results on the gel are then “transferred” to a membrane that can be analyzed more easily. Typically, this is a sheet of nitrocellulose. A blocking agent is then applied to the membrane to prevent nonspecific binding of the “primary antibody”. The primary antibody’s job is to latch onto the proteins that were transferred to the membrane from the gel. This antibody is specifically picked to bind to the target proteins. In order to create a readable pattern on the membrane, a “secondary antibody” is added. This attaches to the primary antibody, and will fluoresce under a certain wavelength of light. After a few washes in Tris-buffered Saline in order to rid the membrane of any excess antibodies, a UV scanner (which can expose the membrane to a specific wavelength) is used in order to make the secondary antibodies fluoresce. What is left is a series of “bands” which the gel electrophoresis created. If a scientist is checking to see whether or not a target strand of DNA is being expressed, they can simply check to see if the size of the proteins in their sample matches the weights of the proteins that the DNA is expected to produce. This complicated process performs a simple but invaluable test. While it doesn’t indicate with certainty that a gene is causing a cell to demonstrate a physiological change, it does indicate that the gene is telling the cell what to do. My Senior Project began on Tuesday the 7th, I had just flown into New York and was excited to begin my experience at the Department of Chemistry and Life Sciences at the United States Military Academy. Throughout the next three months, I am looking forward to learning all about bio-engineering, as well as how a real-life lab operates. After this first week, I am happy to say that I am well on my way to doing just that.
For the majority of the past four days, my time in the lab has been spent shadowing Dr. Alex Mitropoulos. Dr. Mitropoulos’s work has centered on the development of an “eNOSE”, a device that can utilize the biological pathways that allow living things to detect odorants in order to identify volatile substances. However, before a physical prototype can be constructed, it is important to remember that the biology behind it must be proven effective. To show that it is possible to engineer a cell to respond to an odorant (and for that response to be monitored and interpreted), a workflow had to be developed for the lab. My first day in the facility was spent learning about the process by which this lab’s research is conducted. Firstly, a strand of DNA that codes for an olfactory receptor is received from another USMA lab. A good amount of this DNA is needed, so the original copy is inserted into bacteria which will readily divide and amplify the genetic material. A small beaker ripe with nutrients is the perfect environment for this to happen, it only takes one night for the solution to turn cloudy with bacteria. Although the target DNA is being produced by the bacteria, it is mammalian cells that will be used as sensors in the eNOSE. As such, the DNA must be extracted from the bacteria in a process referred to as “purification” so that it can be transferred into the mammalian cells (this is referred to as “transfection”). A series of solutions purify the DNA by discarding the unneeded cell membranes, proteins, and RNA strands. Transfection is accomplished by a compound known as “Lipofectamine”. This incredible substance forms micelles around the DNA, mimicking a lysosome that eukaryotes will readily accept. This allows the DNA of interest to “Trojan Horse” its way into the target mammalian cells. Finally, I got some experience in the simple (but very important) task of keeping the mammalian cells alive. While bacteria are hardy, the cells which will act as sensors in the eNOSE are quite needy. Every few days, their media must be replaced in order to provide them with enough nutrients to survive. I am very excited to observe the last steps of the workflow in the coming days and weeks. Specifically, I am interested in seeing the different ways that the success of the transfection can be gauged. From what I have heard, performing a Western Blot can reveal if the DNA is being expressed by showing the production of a target protein, but an analysis of the cells’ electrical activity in the presence of an odorant will prove the existence of the desired physiological response. After this successful week, my motivation to keep learning and eventually take up a more active role in the lab can only increase. Thank you for reading the first installment in my blog, feel free to comment below if you have any feedback! |
AuthorIsaac Dabkowski - 2017 BASIS Oro Valley senior Archives |
 RSS Feed
RSS Feed
